Spark™ delivers a broader range of accessible chemistries than ever before with the addition of SureChEMBL fragment databases
New and updated fragment databases from ChEMBL and SureChEMBL are now available in Spark. This new release of databases significantly expands the available chemistry to Spark users by including the latest compounds from scientific literature, as well as previously unseen compounds from chemical patents. Combined with databases of fragments derived from screening compounds, and with custom databases which you can generate from your corporate collections using the Spark Database Generator, they provide an outstanding source of bioisosteres you can use to generate new ideas for your project.
More than 1.6 million fragments from ChEMBL
The updated Spark ‘ChEMBL’ databases include an expanded choice of more than 1.6 million fragments derived from release 30 of ChEMBL, a collection of around 2.1 million compounds reported in peer-reviewed scientific literature.
More than 14 million fragments derived from SureChEMBL
In addition, new ‘SureChEMBL’ fragment databases are available to Spark users for the first time. These include more than 14 million fragments derived from the SureChEMBL collection of approximately 17 million compounds harvested from the patent literature. This addition affords Spark users a significant expansion on fragment databases derived from ChEMBL, providing a broader range of accessible chemistries than ever before.
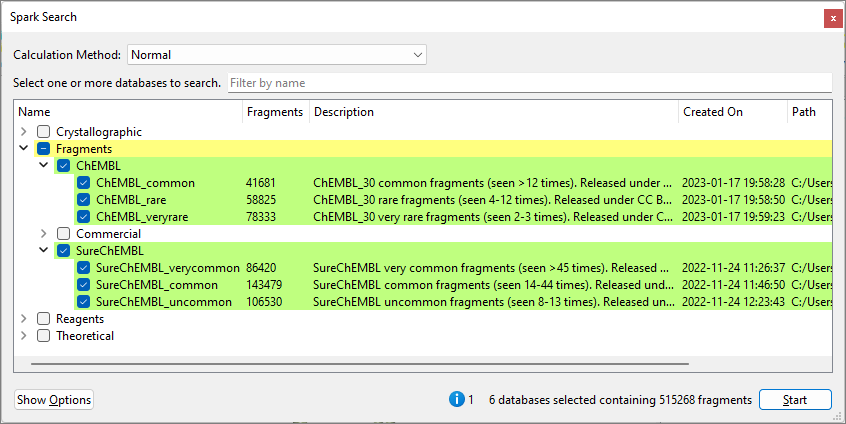
Figure 1. Spark Search window for an R-group replacement experiment, showing default ChEMBL and SureChEMBL fragment collections with a single attachment point.
Like ChEMBL, SureChEMBL compounds come with property information such as synthetic tractability, toxicity, and metabolic reactivity. Compounds in the original source collections were filtered on these biochemical properties prior to fragmentation in Spark, removing molecules containing potentially toxic or reactive groups. Compounds were then cleaved by breaking the bonds connecting carbon atoms to functional groups such as heteroatoms, carbonyls, thiocarbonyls and rings, whilst preserving functional groups such as carboxylic acids, nitro groups and rings. All fragments were subject to heavy atom count and rotatable bond limits, which increases the probability that fragments will form part of biochemically active small molecules. Finally, fragments were sorted by frequency of occurrence in the source dataset and grouped by commonality to form Spark databases. A key assumption for Spark users is that regularly occurring fragments are likely to form part of novel active biochemistry, therefore we recommend that users try the more common libraries before moving on to less common.
Table 1 reports the total number of fragments in each Spark database, with the frequency of occurrence in the source dataset. Database frequencies were arbitrarily chosen to give manageable database file sizes.
Table 1. Fragment databases sorted by frequency.
| Spark category | Database | Total number of fragments (x1000) | Frequency in source dataset |
| ChEMBL | Common | 223 | Fragments which appear in more than 12 molecules |
| Rare | 304 | Fragments which appear in 4-12 molecules | |
| Very Rare | 390 | Fragments which appear in 2-3 molecules | |
| Extremely Rare* | 783 | Fragments which appear in 1 molecule | |
| SureChEMBL | Very Common | 509 | Fragments which appear at least 45 molecules |
| Common | 795 | Fragments which appear in 14-44 molecules | |
| Uncommon | 554 | Fragments which appear in 8-13 molecules | |
| Rare* | 957 | Fragments which appear in 5-7 | |
| Very Rare* | 757 | Fragments which appear in 4 molecules | |
| Extremely rare* | 979 | Fragments which appear in 3 molecules | |
| Doubleton* | 2,545 | Fragments which appear in 2 molecules | |
| Singleton* | 4,794 | Fragments which appear in 1 molecule |
*Contact us for further details.
The number of compounds per connection point count in each database is presented in Figure 2.
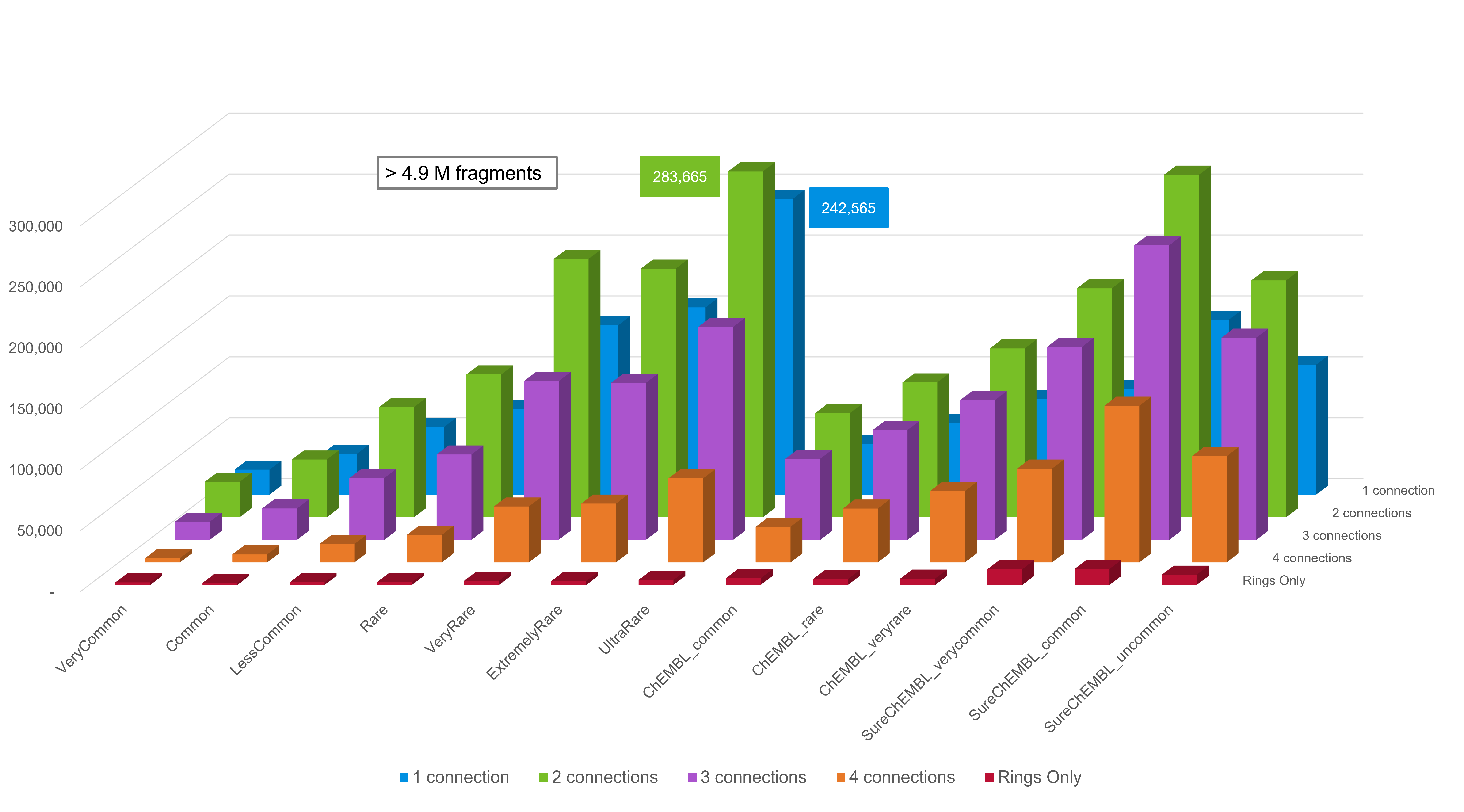
We recommend that users only install ChEMBL databases where the fragment frequency is at least 2, and the three SureChEMBL ‘Common’ databases (Very Common, Common and Uncommon), as the size of the SureChEMBL databases is very large. Furthermore, singleton and doubleton databases for both ChEMBL and SureChEMBL may contain fragments derived from erroneous structures in the source dataset.
Though ChEMBL and SureChEMBL fragments are from different sources (scientific literature vs. patents), as expected, there is significant overlap between some of the databases, as shown in Table 2. In particular, the ChEMBL Common and SureChEMBL Very Common databases show the highest overlap, which is to be expected because most bioactive compounds contain basic subunits, e.g., phenol or pyridine rings.
Table 2. Overlap of the most common fragments in the ChEMBL and SureChEMBL databases.
| SureChEMBL | ||||||||||
| Very Common | Common | Un-common | Rare | Very Rare | Extremely Rare | Doubleton | Singleton | Unique | ||
| ChEMBL | Common | 66% | 14% | 3% | 3% | 1% | 1% | 2% | 2% | 8% |
| Rare | 28% | 21% | 7% | 7% | 3% | 3% | 4% | 5% | 23% | |
| Very Rare | 13% | 17% | 7% | 8% | 4% | 5% | 7% | 6% | 33% | |
| Extremely Rare | 6% | 10% | 6% | 7% | 4% | 5% | 8% | 13% | 42% | |
Reagent databases
The January update of the Spark reagent databases includes around 293,000 reagents derived from the eMolecules building blocks using an enhanced set of rules for chemical transformation. These databases are updated monthly to give you up-to-date availability information, to make it easy for you to order the reagents you require to synthesize your favorite Spark results.
Update your Spark databases
The updated ChEMBL and new SureChEMBL databases, combined with the Spark Commercial fragment databases, provide more that 19 million unique fragments to search, with more that 4.9 million fragments in the recommended databases alone. They significantly expand the choice of fragments for your Spark experiments, providing an even better source of novel ideals for your drug discovery projects.
Spark users can contact our Support team to update their ChEMBL and SureChEMBL databases.
If you are a medicinal chemist or computational chemist and not currently using Spark, contact us to find out how it can help you generate innovative ideas, explore chemical space and escape IP and toxicity traps, or request an evaluation to try Spark on your project.